RNA Design & Programmable Assembly
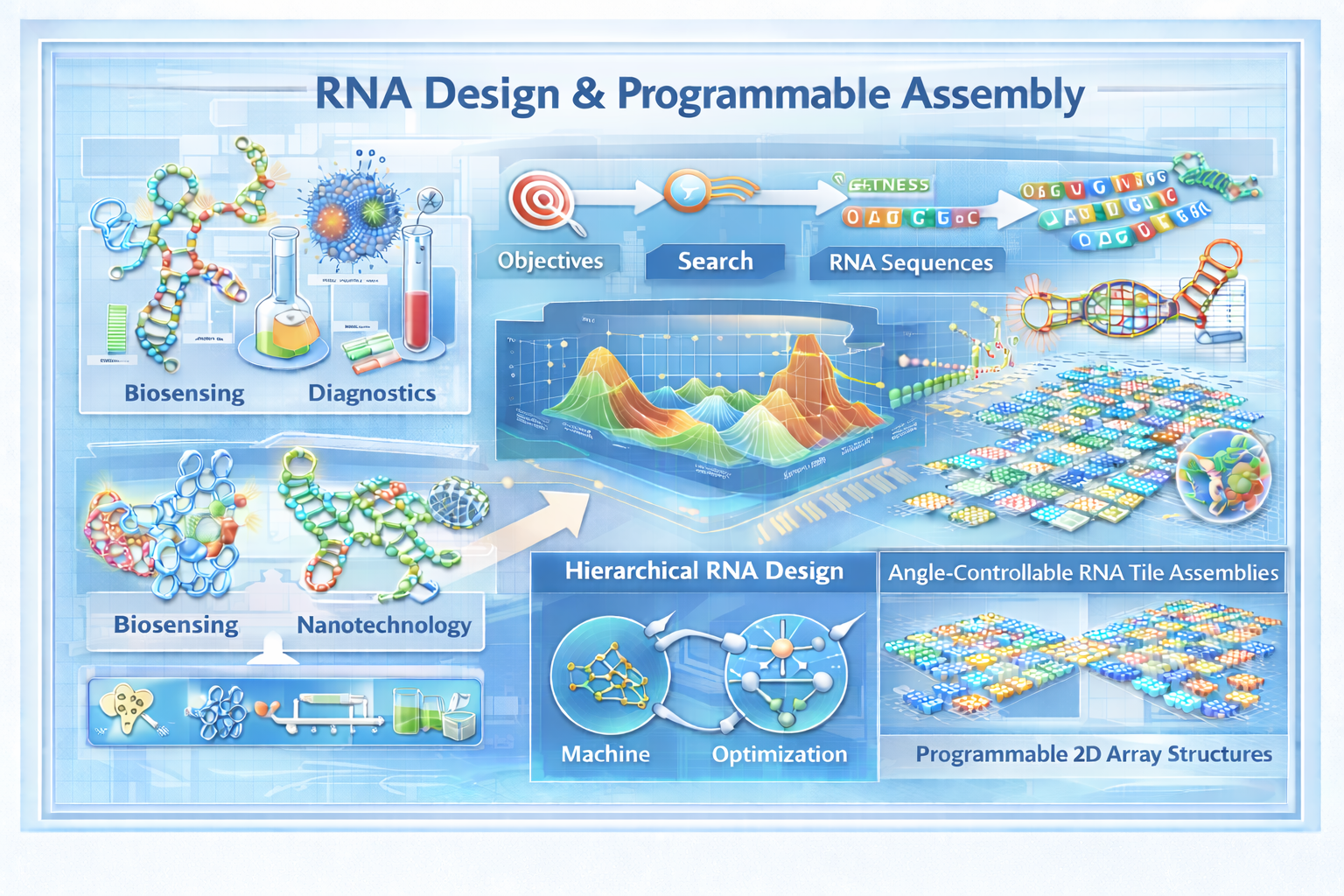
RNA molecules can be rationally engineered to fold into prescribed three-dimensional shapes, enabling applications in biosensing, diagnostics, therapeutics, and programmable nanotechnology. The challenge is a complex inverse design problem: given a desired structure or function, find the sequence — or sequences — that achieve it. The search space is astronomically large and the fitness landscape highly non-convex, making this a natural target for advanced black-box optimization.
Our group develops hierarchical computational frameworks that integrate structure-aware machine learning, optimization, and experimental feedback to navigate this landscape efficiently. In recent work, we demonstrated angle-controllable RNA tile assemblies that enable programmable two-dimensional array structures, published in Nature Communications. Alongside this, we have developed deep learning models that predict RNA sequence-structure likelihood, providing a fast surrogate for guiding the search toward structurally plausible candidates.
Representative Publications
- Yang, Q., Chang, X., Lee, J.Y., Wisniewski, H., Zhou, Y., Pedrielli, G., & Zhang, F. Angle-controllable RNA tiles for programmable array assembly and RNA sensing. Nature Communications, 16(1), p.3728, 2025. DOI
- Zhou, Y., Pedrielli, G., Zhang, F., & Wu, T. Predicting RNA sequence-structure likelihood via structure-aware deep learning. BMC Bioinformatics, 25(1), p.316, 2024. DOI
Biomanufacturing
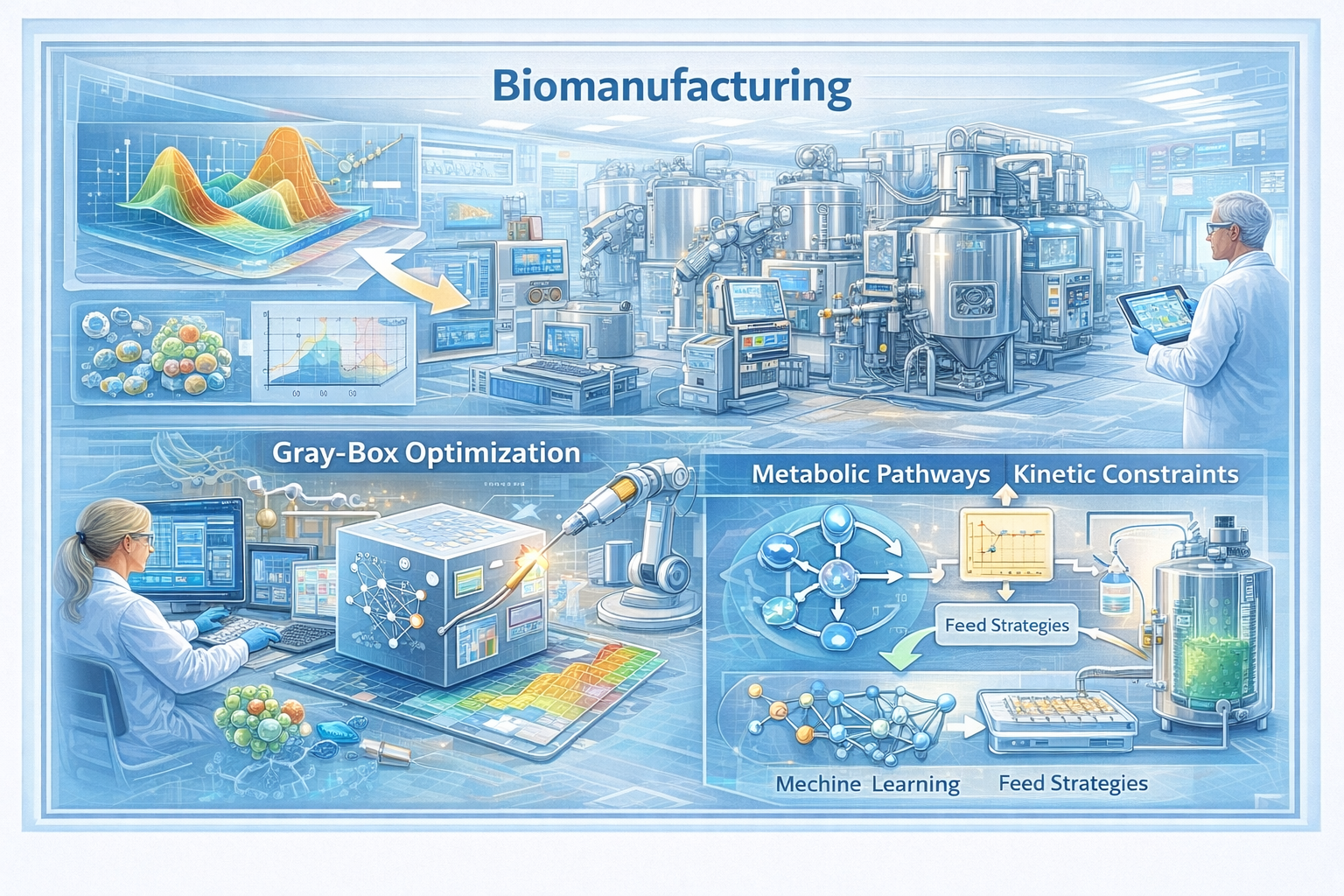
Biomanufacturing — the production of biologics, therapeutic proteins, and cell-based products — involves highly complex bioprocesses that are expensive to run and difficult to model from first principles. Finding optimal operating conditions (media composition, feed strategies, temperature profiles) requires navigating a high-dimensional space with costly experiments and partially understood mechanisms.
We develop gray-box optimization algorithms that exploit partial mechanistic knowledge of biological processes — such as known metabolic pathways or kinetic constraints — to build more informative surrogates and guide the search more efficiently than pure black-box methods. This structure-aware approach reduces the number of experiments needed to reach high-performance operating points, directly supporting the scale-up and cost reduction of biologic production. This line of work is a core application driving our NSF CAREER award on LEarning to Search with Structure (LESS).